


Augmented Tangible Molecular Models main page
Augmented Tangible Molecular Models
Description
Physical models superimposed with virtual computational models and labels using augmented reality. Python extension of ARToolkit in PMV is used for tracking and registering virtual models. Fiducial square markers are used to track and register in three dimensional space via video segmentation. Animations can be used to effectively describe complex structural information about molecular properties.
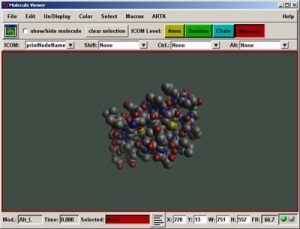
Python Molecular Viewer - PMV
PMV is a Python-based molecular visualization environment developed by the Molecular Graphics Laboratory at The Scripps Research Institute.The new PMV version is written in python 2.3, supports multi-threading and also works on most platforms. Allows viewing of molecular structures from formats such as PDB - PDBQ , PDBQS Auto Dock formats. PQR Mead & Mol2 Tripos format.It Supports multiple representations of molecular structure.
Documentation for PMV is available in PDF form.
Multi-modal user interface for teaching structural molecular biology.
- Complex physical models developed with latest automated model fabrication technology.
- Virtual models overlaid on the physical models using augmented reality.
- Haptic feedback for feeling the electrostatic force fields surrounding physical models in augmented reality.
- Integrated voice module for “hands-free” interactions.
- Augmented animations for better understanding of molecular properties.
- Comprehensive Help Menu system providing real-time interaction for students.

The Protein Structure Magic Book
Proteins have several levels of structural organization: primary, secondary, tertiary, and quaternary. A protein’s primary structure is the sequence of its amino acids. The forces between these amino acids cause the chain to fold and to form the protein’s secondary, tertiary, and quaternary structures. In this book, the characteristics of each level of structure are described and illustrated with a 3-D animation.
There are three Coloring Schemes to represent the protein structures and amino acids used in this book; Color by Atom Type, Color by Amino Acid Type and Color by Chain.
Contacts
Suzanne Weghorst <weghorst![]() u.washington.edu>
u.washington.edu>